min_spanning_tree#
- graph_tool.topology.min_spanning_tree(g, weights=None, root=None, tree_map=None)[source]#
Return the minimum spanning tree of a given graph.
- Parameters:
- g
Graph Graph to be used.
- weights
EdgePropertyMap(optional, default: None) The edge weights. If provided, the minimum spanning tree will minimize the edge weights.
- root
Vertex(optional, default: None) Root of the minimum spanning tree. If this is provided, Prim’s algorithm is used. Otherwise, Kruskal’s algorithm is used.
- tree_map
EdgePropertyMap(optional, default: None) If provided, the edge tree map will be written in this property map.
- g
- Returns:
- tree_map
EdgePropertyMap Edge property map with mark the tree edges: 1 for tree edge, 0 otherwise.
- tree_map
Notes
The algorithm runs with \(O(E\log E)\) complexity, or \(O(E\log V)\) if root is specified.
References
[kruskal-shortest-1956]J. B. Kruskal. “On the shortest spanning subtree of a graph and the traveling salesman problem”, In Proceedings of the American Mathematical Society, volume 7, pages 48-50, 1956. DOI: 10.1090/S0002-9939-1956-0078686-7 [sci-hub, @tor]
[prim-shortest-1957]R. Prim. “Shortest connection networks and some generalizations”, Bell System Technical Journal, 36:1389-1401, 1957.
Examples
>>> from numpy.random import random >>> g, pos = gt.triangulation(random((400, 2)) * 10, type="delaunay") >>> weight = g.new_edge_property("double") >>> for e in g.edges(): ... weight[e] = linalg.norm(pos[e.target()].a - pos[e.source()].a) >>> tree = gt.min_spanning_tree(g, weights=weight) >>> gt.graph_draw(g, pos=pos, output="triang_orig.pdf") <...> >>> u = gt.GraphView(g, efilt=tree) >>> gt.graph_draw(u, pos=pos, output="triang_min_span_tree.pdf") <...>
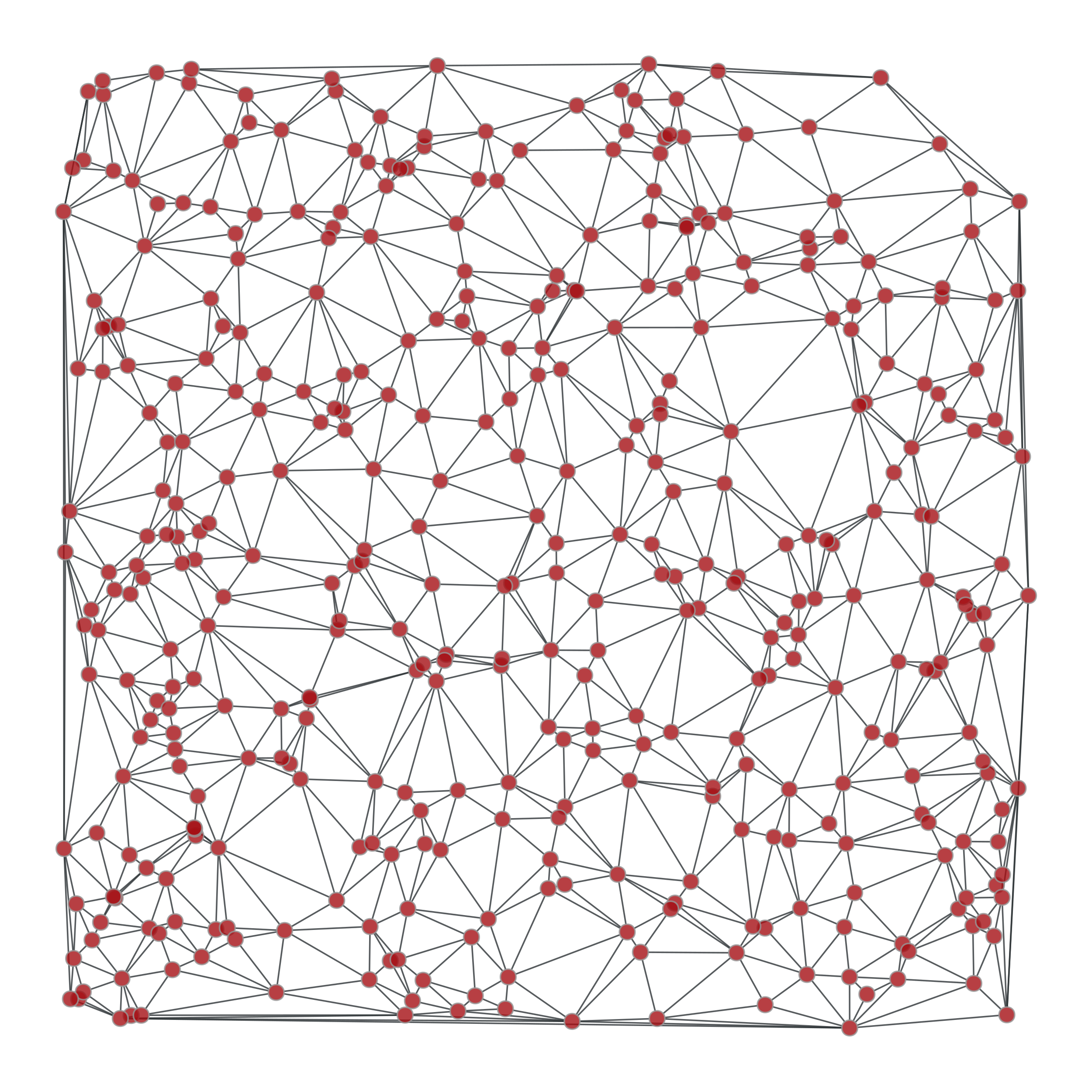
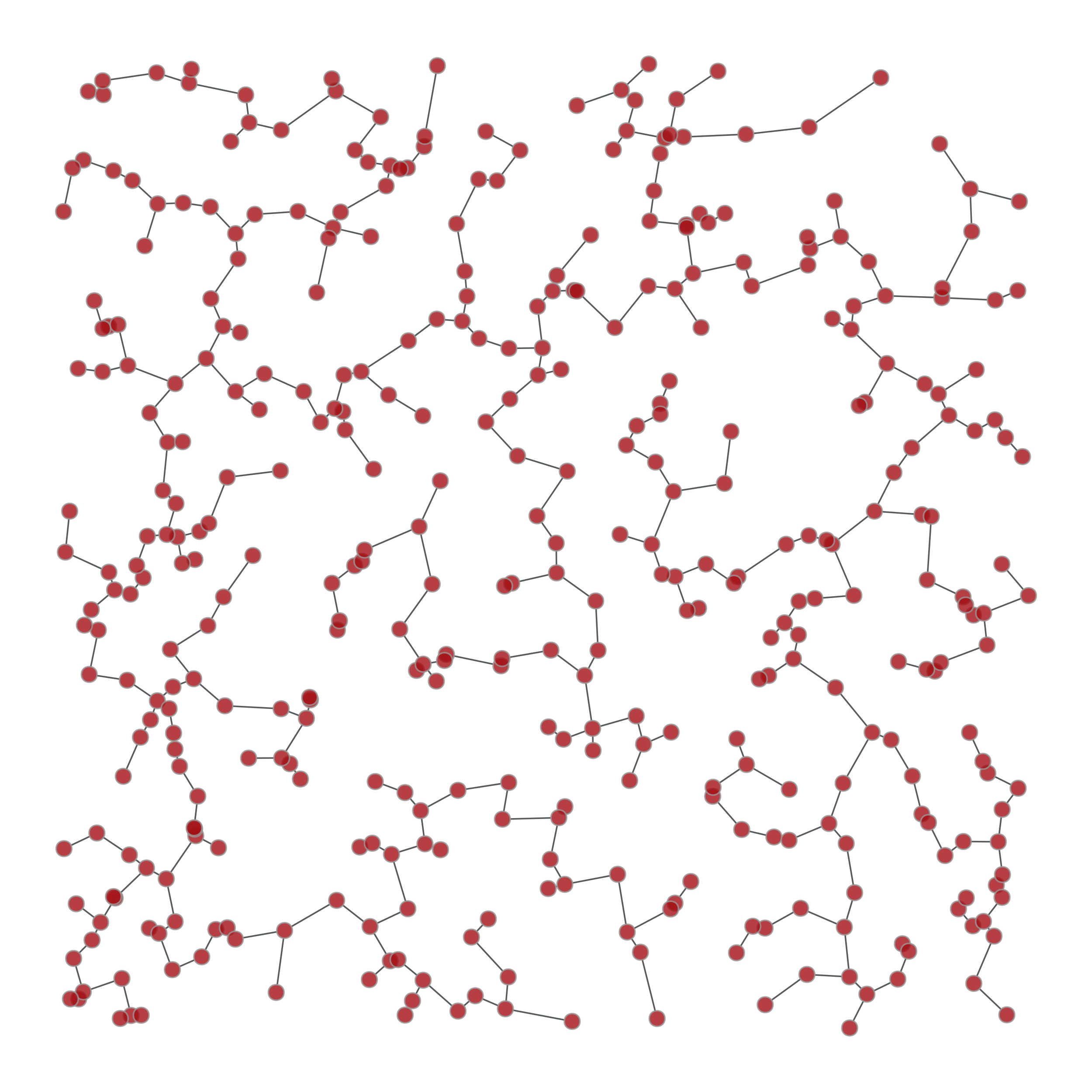
Left: Original graph, Right: The minimum spanning tree.